ASKAPsoft processing of ASKAP data¶
Introduction¶
ASKAPsoft is the suite of processing software developed by the ASKAP computing team to handle the pipeline processing of ASKAP observations. It has been primarily designed for full-scale ASKAP processing on a high-performance computing environment. The design philosophy and background is detailed in the ASKAP Science Processing document (ASKAP-SW-0020), which was produced in 2011 to detail the entire ASKAP pipeline processing environment. The current processing uses a slightly different approach, particularly in the calibration method, and the set of scripts detailed here are designed to demonstrate a reduction approach for ASKAP with ASKAPsoft.
This set of scripts handles all reduction steps, from the splitting and flagging of bandpass-calibrator and science datasets, obtaining and applying the bandpass calibration, averaging of the science visibilities, continuum imaging (with or without self-calibration) and linear mosaicking, source-finding, and spectral-line imaging (including continuum subtraction and gains calibration).
The scripts are bash shell scripts, which create parsets and slurm files that are submitted to the queue on galaxy. The jobs have their dependencies set appropriately, so that a single call of the control script is sufficient to run all desired processing.
The scripts have been designed to be controlled by a single user
input, wherein most parameters can be set. The following tables detail
the parameters that are available to be set by the user. It is also
possible to manually alter the slurm scripts to change parameters that
have not been defined for user input (the slurm files can be seen in
the slurmFiles/ directory - if you set SUBMIT_JOBS=false, they
will be created but not submitted). If there are parameters that you
would like defined for user input that are currently hardcoded, please
let me know.
Finally, note that the processing must not be run on /home. The /home filesystem is not suitable for running this processing, and there are certain bits that will fail on /home. If the filesystem on which the scripts are run is not a Lustre filesystem, it will exit without running anything. This is because the Lustre striping is set at run-time (to 4 for the processing directories, and 1 for other directories like parsets, logs & metadata).
Assumptions¶
These scripts have been put together to process particular observations. The assumed structure of your observation is that you have an observation of a ‘science’ field (i.e. the field for which you want to make images - this can actually incorporate more than one FIELD location, through interleaving), and an observation of 1934-638 which will be used to determine the bandpass solution. This should have been done in the usual calibration way of short observations of 1934-638 in each beam sequentially.
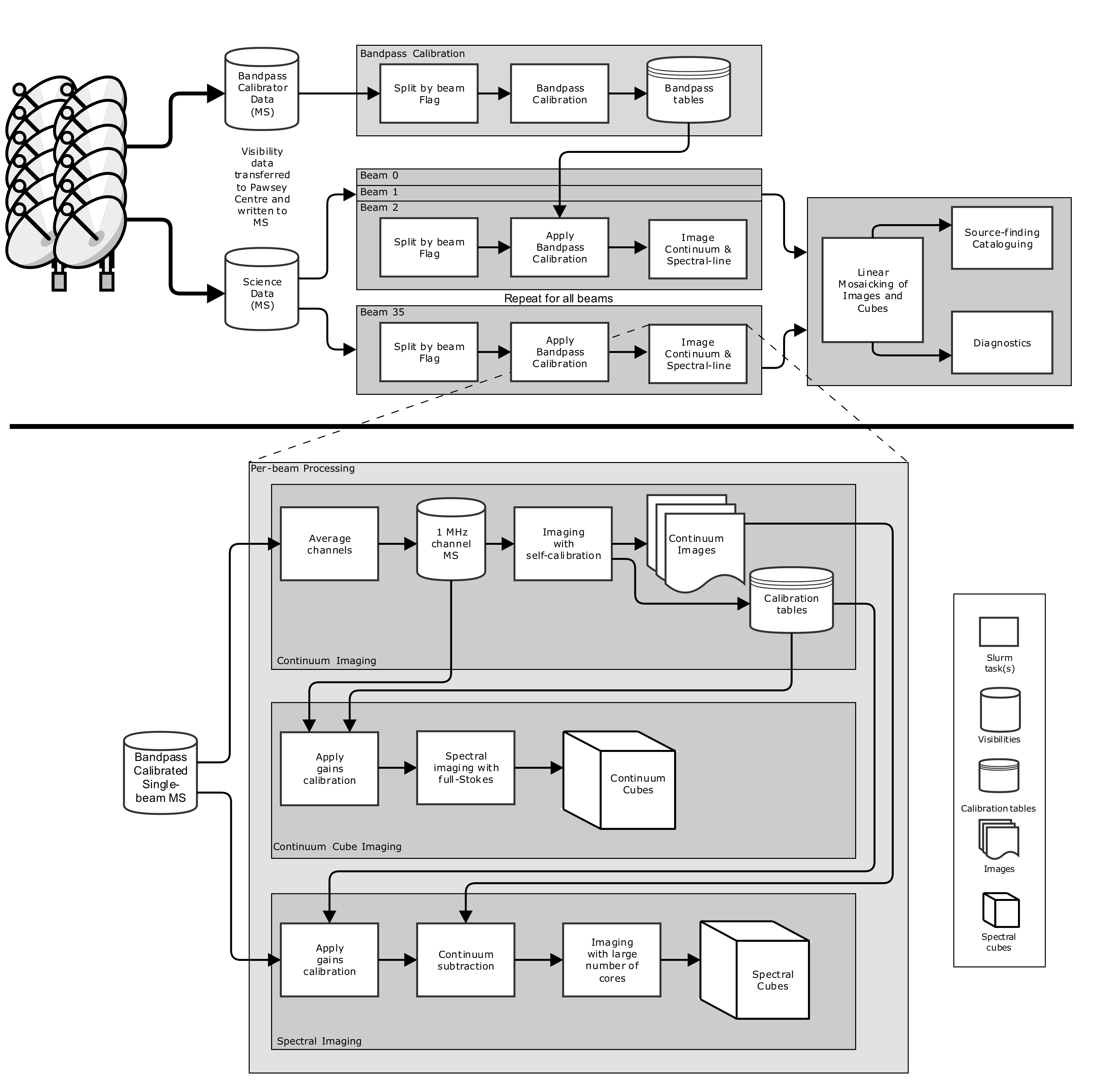
Overview¶
The following diagram indicates the overall flow of processing. Details of each step can be found in How to run the ASKAP pipelines, and in the specific pages on each section.