Solvers¶
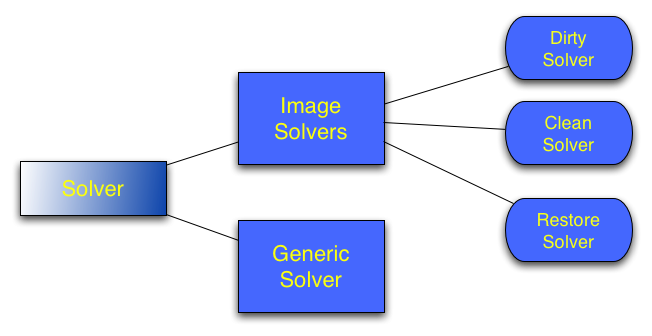
A number of solvers are available in ASKAPsoft. For imaging purposes only specialized image solvers are generally used, although generic SVD-based linear solver (see the diagram above) can be used with (very small) images as well (some change to the code may be required as this is not a normal use case). The solver parset parameter defines the type of the image solver to use with the choice between Dirty, and Clean. The Dirty solver just inverts the data (takes the normal equations and simply divides the data vector by the diagonal of the normal matrix. This is analogous to making the dirty image or a linear mosaic of dirty images), while the Clean performs minor cycle cleaning. If multiple beams and/or fields are present in the dataset (and mosaicing gridders (see Gridders for details) are used), joint deconvolution is always preformed (individual deconvolution is not supported).
The Restore solver is somewhat special. It is executed behind the scene with very minimal setup required from the user (the same parameters are generally used as for the cleaning solver). Restore solver is described separately in the cimager.
Common Parameters¶
Parameters for all solvers:
Parameter |
Type |
Default |
Description |
|---|---|---|---|
solver |
string |
none |
Selection of solver. Specify either “Clean” or “Dirty” |
verbose |
string |
“true” |
True enables lots of output |
tolerance |
float |
0.1 |
cutoff value given as a fraction of the largest diagonal element. The linear system describing interferometric measurement is inverted approximately, assuming that the matrix is diagonal, i.e. the right-hand side is divided by the appropriate diagonal element (which is a weight). If diagonal is smaller than this tolerance multiplied by the largest diagonal element, the right-hand side instead is either divided by the largest diagonal element to get the result or the result is set to zero. This is controlled by weightcutoff parameter. For images it means that areas with low weight (i.e. a mosaic edge) are not boosted up. In addition to weight truncation, all pixels with the weight below cutoff are normally masked out. The weightcutoff.clean parameter allows to assign mask corresponding to weight truncation. This allows S/N-based cleaning to happen, if the peak of S/N is realised outside the nominal field of view. |
weightcutoff |
string |
truncate |
Either “zero” or “truncate”. This parameter controls what actually happens for values below cutoff defined by the tolerance parameter. If zero is given, the appropriate values are set to zero. For truncate, the values are divided by the largest diagonal. |
weightcutoff.clean |
bool |
false |
This parameter defines whether the values below cutoff are masked out or not. By default, the are masked out and so S/N-based clean never finds optima among these values. If this parameter is true, the mask is actually sqrt(tolerance), which corresponds to truncation of the diagonal during normalisation. This potentially allows cleaning to happen, if no peak of the S/N is realised among these values. |
Parameters verbose and tolerance have solver.Clean or solver.Dirty prefix. Although verbose is understood by dirty solver, there is currently no effect. Dirty solver doesn’t require any additional parameters (but one can also set up a preconditioner described in the following section with the dirty solver). Additional parameters understood by the Clean solver are given in the following section.
Clean Solver Parameters¶
All parameters given in the next table have solver.Clean prefix (i.e. Cimager.solver.Clean.algorithm) or Cimager.solver.Clean.scales. The Clean solver with algorithm set to “Basisfunction” is an improved version of the casacore LatticeCleaner. Most importantly, it supports use of a patch in the deconvolution. This decreases memory use and run time by approximately the ratio of pixels in the patch to pixels in the image.
Parameter |
Type |
Default |
Description |
|---|---|---|---|
algorithm |
string |
“BasisfunctionMFS” |
Valid choices are “MultiScale”, “Basisfunction”, “Hogbom”, “MultiScaleMFS” and “BasisfunctionMFS”. Use “Hogbom” for a single scale, non-MFS case. For the Clean solver, the casacore’s LatticeCleaner used to do the actual work in non-MSMFS case will be set up with CleanEnums::HOGBOM if algorithm is Hogbom and the single scale of 0 will be used. For the Basisfunction algorithm, a re-implemented and improved version of the CASA MultiScale algorithm is used. BasisfunctionMFS is the optimised algorithm used for most ASKAP processing |
scales |
vector<string> |
[0, 3, 10] |
Scales to be solved (defined in pixels). Ignored if algorithm=”Hogbom” |
niter |
int |
100 |
Number of minor cycles |
gain |
float |
0.1 |
Loop gain. Fraction of the peak subtracted during one minor cycle. |
speedup |
float |
no speed up |
Relevant for “MultiScale” and “MultiScaleMFS” only. If defined, the value will be passed as a speed up factor to the lattice cleaners doing the minor cycle. According to casacore’s manual, this will speed up clean by raising the threshold (could help if the threshold set too low for the given dataset). The physical meaning of the parameter is the number of iterations required to double the threshold. |
padding |
float |
1.0 |
Optional padding of all images in the solver (minor cycle will be done on an image this factor times larger on both axes, e.g. to alleviate the fact that FFT is used to compute convolutions). Default value means no padding. At this stage this option is understood by MultiScale cleaner only |
logevery |
int |
1 |
How frequently to log progress in the minor cycle. Every nth iteration is reported (ie. if logevery=100, every 100th iteration is reported), providing the iteration number, the peak residual, the objective function and the total flux. |
saveintermediate |
bool |
true |
Save intermediate images (residuals and preconditioned PSF) at the end of each majorcycle. |
The following parameters are available for the Basisfunction and BasisfunctionMFS algorithms.
Parameter |
Type |
Default |
Description |
|---|---|---|---|
psfwidth |
int |
0 |
Sets the width of the psf patch used in the minor cycle. This decreases memory use and run time by approximately the ratio of pixels in the patch to pixels in the image. |
detectdivergence |
bool |
true |
Check if the deconvolution is diverging - stop the major cycles if the residuals increase by a factor 2 |
detectmilddivergence |
bool |
false |
Check if the deconvolution is starting to diverge. Go to the next major cycle if residuals increase by 10% from the lowest residual found this cycle, but only if more than 25% of the iterations have been completed. |
divergencelevels |
vector<float> |
|
Set the parameters for divergence detection: 1st - mild minor cycle divergence, increase from smallest minor cycle residual, 2nd - fatal minor cycle divergence, increase from starting residual 3rd - fatal major cycle divergence, increase from previous starting residual 4th - fraction of iterations to complete before mild divergence detection is turned on |
orthogonal |
bool |
false |
Use orthogonal spatial basisfunctions - this does not appear to be as useful as expected - avoid in normal use |
usecleanmask |
bool |
false |
(BasisFunctionMFS) If true look for user specified clean mask image(s). They should have the same naming scheme as other images but start with ‘cleanmask’ and be of float type with 1.0 for included and 0.0 for excluded pixels. |
useoverlapmask |
bool |
true |
(BasisFunctionMFS) Use a mask on the main image to exclude areas covered by offset fields. May need to turn this off when faceting. |
usepixellists |
bool |
true |
(BasisFunctionMFS) Use a list of pixels above the clean threshold to do peak searching and residual cleaning instead of sarching and cleaning the full images |
usepixellists.tolerance |
float |
0.1 |
(BasisFunctionMFS) Fraction to lower the threshold by to include pixels in the list that may be boosted above the threshold while removing negative sidelobes. Try 0.1 or 0.2, 0.5 and above may cause divergence |
usepixellists.nsigma |
float |
4.0 |
(BasisFunctionMFS) Use a noise threshold for the residual image at each scale to avoid overcleaning some scales leading to loss of flux. This also avoids getting too many pixels in the pixel list and slowing progress. However, if the residual image has emission in more than 50% of the pixels set this to 0.0 as the noise estimate will fail in that case and not much will be cleaned. |
usepixellists.npixrange |
vector<int> |
[10,100] |
(BasisFunctionMFS) The default specifies that the maximim number of pixels collected for peak searching is to be within 10 to 100 times (default) the number of iterations. This default works well for fields dominated by point sources and small extended sources. For fields dominated by large scale structure (e.g. GASKAP survey) set this higher [50,200] to achieve results similar to the non-pixellist clean. Note that too many pixels in the pixel list slows progress, so the default is lower. For mostly empty images (point sources) you can set this lower, e.g., [2,10]. An alternative to increasing this is to turn on useincrements - it also helps recover extended flux |
useincrements |
bool |
false |
(BasisFunctionMFS) skip pixels at larger scales. When using large scales, nearby pixels will be strongly strongly correlated. This feature on skips progressively more pixels in the peak search as the scale size grows. For pixellists this helps limit the size of the list. Set to false by default, because it may increase the risk of clean divergence. |
scalemask |
string |
“” |
(BasisFunctionMFS) Read in an initial 2D scale mask with the image name given. The clean will look for components at each scale only where the corresponding scale mask bit is set. Mask image size must be equal or larger than other images (central part will be used), cellsize, direction and scales should match the other inputs. Default is not to read a scalemask. |
noiseboxsize |
uint |
0 |
(BasisFunctionMFS) Set the size of the box to calculate local noise statistics for the noise thresholds. The default of 0 will use the overall image noise level |
positivity |
vector<bool> |
[false] |
(BasisFunctionMFS) For each scale specify if components should be constrained to be positive. Default is false for all scales. Constraining the largest scales to be positive may help keep large negative bowls from being included in the model |
scalebias |
float |
1.0 |
(BasisFunctionMFS) The scalebias parameter, valid range is 0 to 2, best range to try is 0.7 to 1.3. Values less than 1 will prefer large scales in the peak search, values greater than 1 favour small scales. The default of 1.0 works for most cases, but if you find spurious large scale components in the model or image, try increasing this to e.g., 1.1. The factor for each scale is bias^(1+log2(scale/scale1)) where scale1 is the first non zero scale. |
All parameters given in the next table do not have solver.Clean prefix (i.e. Cimager.threshold.minorcycle).
Parameter |
Type |
Default |
Description |
|---|---|---|---|
threshold.minorcycle |
vector<string> |
no threshold |
If defined, the parameter can be either a single string or a vector of 2 or 3 strings. A number without units, or with a percentage sign, is interpreted as a fractional stopping threshold (with respect to the peak residual). An absolute flux given in Jy or related units is interpreted as an absolute threshold. A string like 5.0sigma will set a threshold relative to the noise in the residuals. You can mix fractional and absolute or noise thresholds, but can’t mix absolute and noise ones. An undefined parameter means no minor cycle thresholding is done. A second absolute flux (or noise) threshold can be used to specify a deep clean threshold. During deep cleaning only pixels already in the model are searched to find new components. Setting the deep clean threshold to 0.5 times the noise level generally leaves very few sidelobes visible. Deep clean and noise based threshold are currently only implemented for the BasisFunctionMFS solver |
threshold.majorcycle |
string |
-1Jy |
The target peak residual. Use negative value to ensure all requested major cycles are done. |
threshold.firstimage |
bool |
true |
In case of multiple images (e.g., main + offset) treat subsequent images as if they’re part of the first image. I.e., clean can’t go deeper than it did in the first image at each major cycle. |
threshold.masking |
float |
-1 |
If the value is negative (default), a signal-to-noise based cleaning is done. In other words, a peak of S/N is searched at every minor cycle, rather than a flux peak. A positive value reverts the algorithm back to the traditional absolute flux peak-based clean. In this case, the value is the threshold used for masking on the basis of the weight. For example, a value of 0.9 (btw, this is the default in the casacore’s LatticeCleaner, and, therefore, could be implicitly adopted in casa imager) means that all pixels with less than 90% weight (defined as square root from the ratio of matrix diagonal to the maximum diagonal element) will be masked out for cleaning purposes. |
preconditioner.Names |
vector<string> |
empty vector |
List of preconditioners to be applied (in the order they are given in the list). Preconditioners are ASKAPsoft equivalents of visibility weighting (i.e. uniform, robust, natural), which do not require multiple passes over the dataset. Preconditioners can be viewed as operators applied to equation matrix before it is solved. Having the normal matrix as close to the diagonal as possible (a diagonal form is actually assumed during the inversion process) makes the inversion more accurate. By default, no transformation to the normal matrix is done. This is equivalent to the natural weighting. The following preconditioners are currently implemented: Wiener, NormWiener, Robust and GaussianTaper. In addition, the word None is understood as an empty preconditioner which does nothing. Each preconditioner requires a specific set of parameters described in a separate section. These parameters are given after the name of the preconditioner, e.g. preconditioner.Wiener.noisepower (see below) |
preconditioner.name.xxx |
Use this form to define parameter xxx for preconditioner name. Note, this preconditioner will only be instantiated and used if its name appears in the list given in preconditioner.Names. Description of individual parameters are given in a separate section. |
||
preconditioner.preservecf |
bool |
false |
Use a modified PSF to generate any preconditioner that is derived from the uv sampling function (e.g. Wiener and Robust). This option takes a running mean over an approximate nearest-neighbour sampling function with a box width that is proportional to the support size of the gridding kernels. This enables post-gridding density weighting while preserving the gridding convolutions. Note that this is currently only used with the Wiener preconditioner and the WProject gridder. The default is normally false, but will be true if preconditioning is used. |
Parameters of preconditoners¶
The normal matrix can optionally be transformed by a preconditioning operator before equations are solved. This step can regularise the matrix and improve the quality of the solution. It is the ASKAPsoft way of implementing visibility weighting for the PSF (e.g. uniform, robust), and does not require an additional pass over the data. The following preconditioners are currently implemented: Wiener, NormWiener, Robust and GaussianTaper.
A Wiener filter, which is constructed from the PSF, is the preferred preconditioner. This preconditioner is somewhat analogous to robust weighting.
The preconditioners are described below along with the available parameters. The parameters need a preconditioner.preconditionerName prefix, but not the solver prefix (e.g. Cimager.preconditioner.Wiener.noisepower), although technically preconditioning is done in the solver. When defined this way, the same parameters can be reused in the restore solver (described in the cimager documentation). One exception to this is the restore solver itself, which can have an additional preconditioner specified for the final restore step (e.g. Cimager.restore.preconditioner.Wiener.robustness). The additional restored files will have the .alt.restored suffix.
The table below contains the description of individual parameters (names starts with preconditionerName).
Parameter |
Type |
Default |
Description |
|---|---|---|---|
Wiener.noisepower |
float |
none |
If the Wiener filter is defined with noisepower, this exact value of noise power will be used to construct the filter. Optionally, the PSF can be normalised before the filter is constructed (see normalise option – this is a replacement for the NormWiener preconditioner). Note that the Wiener filter must be specified with either noisepower or robustness, and it is recommended that preservecf is set to true. |
Wiener.normalise |
bool |
false |
This is an additional option for a Wiener preconditioner being constructed from an explicit value of the noise power (i.e. noisepower). If set to true, the PSF will be normalised to 1.0 before the filter is constructed, easing interpretation. Note, this option is incompatible with robustness, because in that case the PSF is always normalised). |
Wiener.robustness |
float |
none |
The noise power is derived from the given value of robustness to have roughly the same effect as the analogous parameter in Robust (i.e., -2.0 close to uniform weighting, +2.0 close to natural weighting). Note that the Wiener filter must be specified with either noisepower or robustness. |
Wiener.taper |
float |
none |
If defined, the FFT of the uv sampling function used to generate the Wiener filter (effectively the PSF) will be tapered with a Gaussian. The value of the parameter is the FWHM of the taper in image pixels. Restricting the filter size to approximately that of the primary beam size is of particular importance when imaging over fields that are larger than the primary beam. There is little point to tapering if preservecf = true. |
NormWiener.robustness |
float |
0.0 |
Roughly the same effect as the same parameter in Robust. |
Robust.robustness |
float |
0.0 |
Post-gridding version of robust weighting is applied. It is recommended that preservecf is set to true. |
GaussianTaper |
vector<string> |
None |
A Gaussian taper is applied to the visibilities. The parameter should be either a single string with the FWHM of a circular gaussian taper, or a vector of three strings for an elliptical taper: the major and minor axis FWHM and the position angle. String values may contain units, e.g. [10arcsec,10arcsec,34deg]. If no units are given, radians are assumed. GaussianTaper currently conflicts with different uv-cell sizes for different images. An exception is thrown if such a condition exists. |
GaussianTaper.isPsfSize |
bool |
false |
If true, try to make the final fitted PSF the size given. This uses the restore.beam.cutoff parameter for the fit. A warning will appear in the log if the target PSF size was not achieved. |
GaussianTaper.tolerance |
double |
0.005 |
Specify the fractional tolerance of the fitted beam size when isPsfSize is true, default is 0.5% |
Examples¶
Multi-scale Clean Solver¶
Cimager.solver = Clean
Cimager.solver.Clean.algorithm = MultiScale
Cimager.solver.Clean.scales = [0, 3, 10]
Cimager.solver.Clean.niter = 10000
Cimager.solver.Clean.gain = 0.1
Cimager.solver.Clean.tolerance = 0.1
Cimager.solver.Clean.verbose = True
Cimager.threshold.minorcycle = [0.27mJy, 10%]
Cimager.threshold.majorcycle = 0.3mJy
Cimager.preconditioner.Names = [Wiener,GaussianTaper]
Cimager.preconditioner.GaussianTaper = [30arcsec, 8arcsec, 10deg]
Cimager.preconditioner.Wiener.noisepower = 100.0
Cimager.ncycles = 5
Cimager.restore = True
Cimager.restore.beam = [30arcsec, 30arcsec, 0deg]
Dirty Solver¶
Cimager.solver = Dirty
Cimager.solver.Dirty.tolerance = 0.1
Cimager.solver.Dirty.verbose = True
Cimager.ncycles = 0